course
Become a ML Scientist
Master Python skills to become a machine learning scientist
Topics
Learn more about Python
4 hours
137.2K
course
Introduction to Python
4 hours
5.5M
course
Intermediate Python
4 hours
1.1M
See More
RelatedSee MoreSee More
tutorial
Python Seaborn Tutorial For Beginners: Start Visualizing Data
This Seaborn tutorial introduces you to the basics of statistical data visualization
Moez Ali
20 min
tutorial
Visualizing Data with Python and Tableau Tutorial
Learn how you can use Python to extend Tableau's data visualization capabilities.
Abid Ali Awan
15 min
tutorial
Introduction to k-Means Clustering with scikit-learn in Python
In this tutorial, learn how to apply k-Means Clustering with scikit-learn in Python
Kevin Babitz
21 min
tutorial
Python Seaborn Line Plot Tutorial: Create Data Visualizations
Discover how to use Seaborn, a popular Python data visualization library, to create and customize line plots in Python.
Elena Kosourova
12 min
tutorial
Altair in Python Tutorial: Data Visualizations
Learn how to create data visualizations in Python using Altair and a Movies Dataset.
Sejal Jaiswal
12 min
tutorial
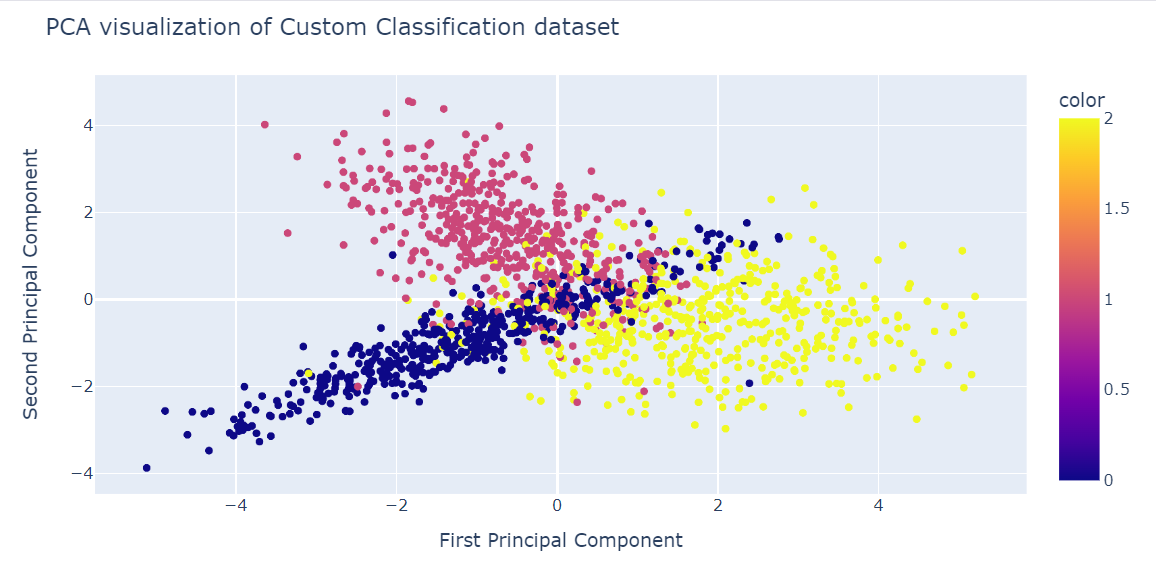
Principal Component Analysis (PCA) in Python Tutorial
Learn about PCA and how it can be leveraged to extract information from the data without any supervision using two popular datasets: Breast Cancer and CIFAR-10.
Aditya Sharma
23 min