track
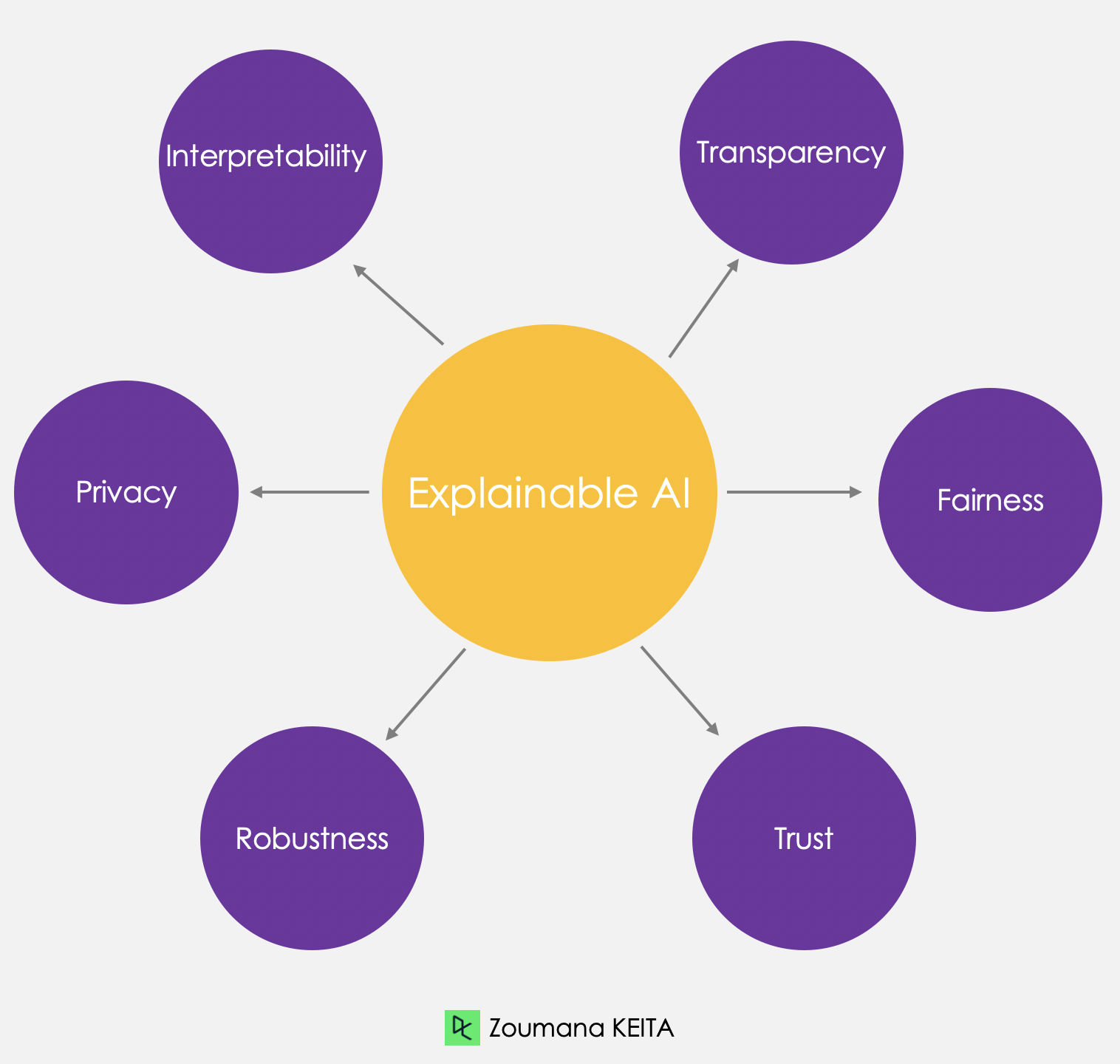
Explainable AI - Understanding and Trusting Machine Learning Models
Dive into Explainable AI (XAI) and learn how to build trust in AI systems with LIME and SHAP for model interpretability. Understand the importance of transparency and fairness in AI-driven decisions.
May 2023 · 12 min read
AI Upskilling for Beginners
Learn the fundamentals of AI and ChatGPT from scratch.
Earn a Top AI Certification
Demonstrate you can effectively and responsibly use AI.
Learn more about AI!
6 hours hours
course
AI Ethics
1 hour
12K
course
Explainable Artificial Intelligence (XAI) Concepts
1 hour
677
See More
RelatedSee MoreSee More
blog
Building trust in AI to accelerate its adoption
Building trust in AI is key towards accelerating the adoption of data science and machine learning in financial services. Learn how to accelerate the development of trusted AI within the industry and beyond.
DataCamp Team
5 min
blog
Understanding and Mitigating Bias in Large Language Models (LLMs)
Dive into a comprehensive walk-through on understanding bias in LLMs, the impact it causes, and how to mitigate it to ensure trust and fairness.
Nisha Arya Ahmed
12 min
blog
What is AI Literacy? A Comprehensive Guide for Beginners
Explore the importance of AI literacy in our AI-driven world. Understand its components, its role in education and business, and how to develop it within organizations.
Matt Crabtree
18 min
blog
What is AI Alignment? Ensuring AI Works for Humanity
Explore AI Alignment: its importance, challenges, and methodologies. Learn how to create AI systems that benefit humanity and align with human values and goals.
Vinod Chugani
12 min
podcast
Interpretable Machine Learning
Serg Masis talks about the different challenges affecting model interpretability in machine learning, how bias can produce harmful outcomes in machine learning systems and the different types of technical and non-technical solutions to tackling bias.
Adel Nehme
51 min
tutorial
An Introduction to SHAP Values and Machine Learning Interpretability
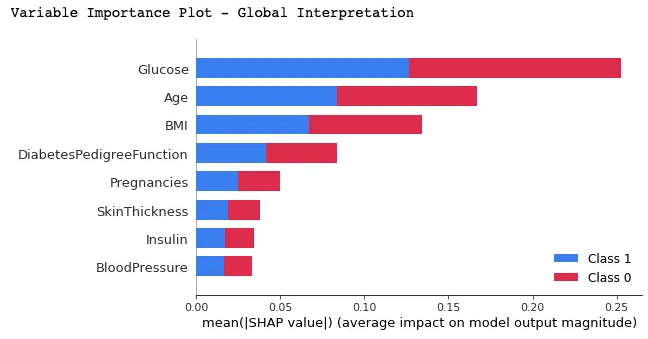
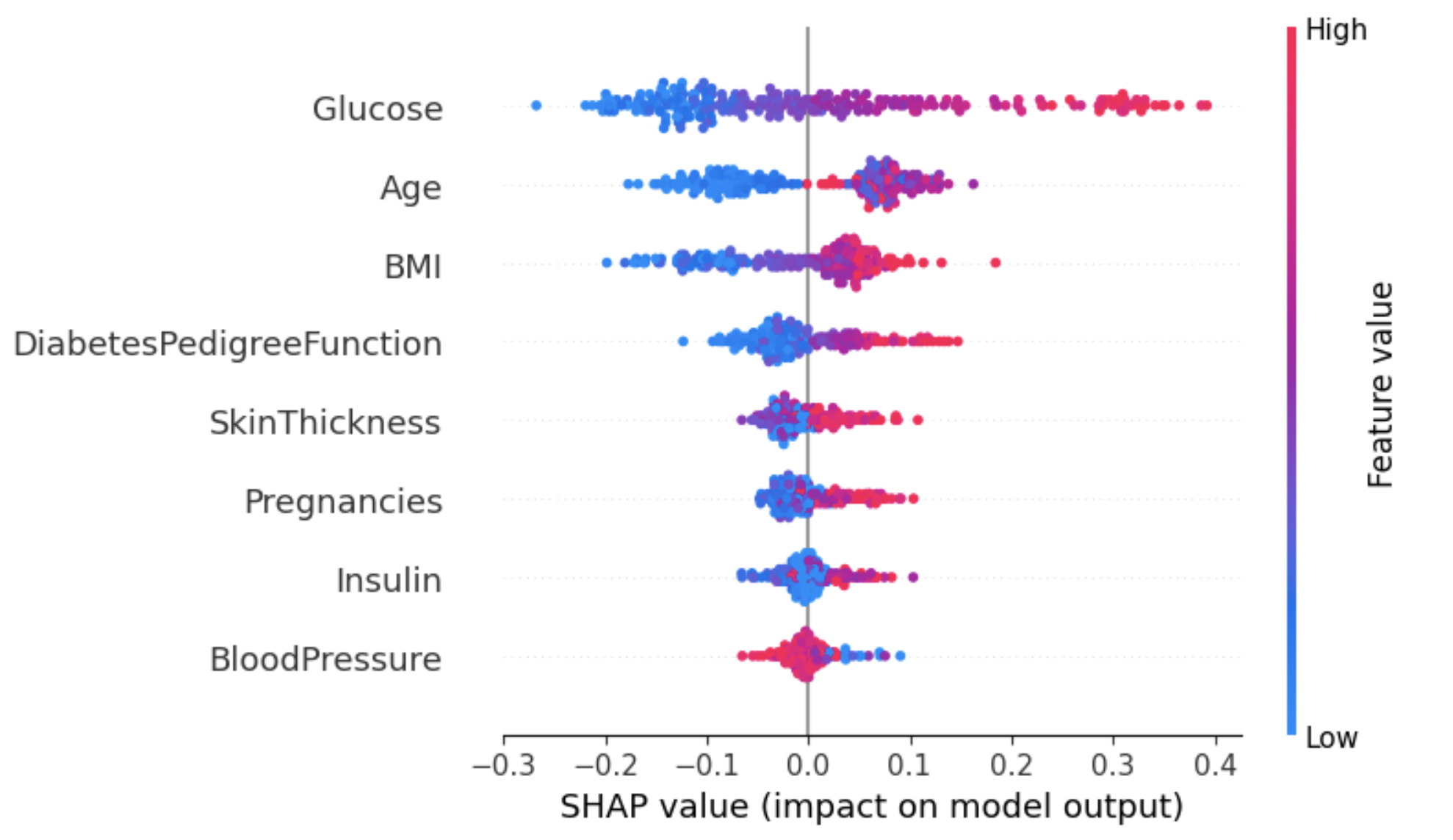
Machine learning models are powerful but hard to interpret. However, SHAP values can help you understand how model features impact predictions.
Abid Ali Awan
9 min